Submit New Annotation
Bystro annotates sequencing data rapidly using multiple databases and provides powerful filtering and natural-language search capabilities. Upload your VCF or SNP files to get comprehensive genomic annotations.
Getting Started
Bystro's submission process supports genomic annotation, data imputation, multi-omics integration, and merging multiple datasets. Follow these steps to submit your data:
Start your submission
Click Submit in the top toolbar and choose your reference genome from the available options.
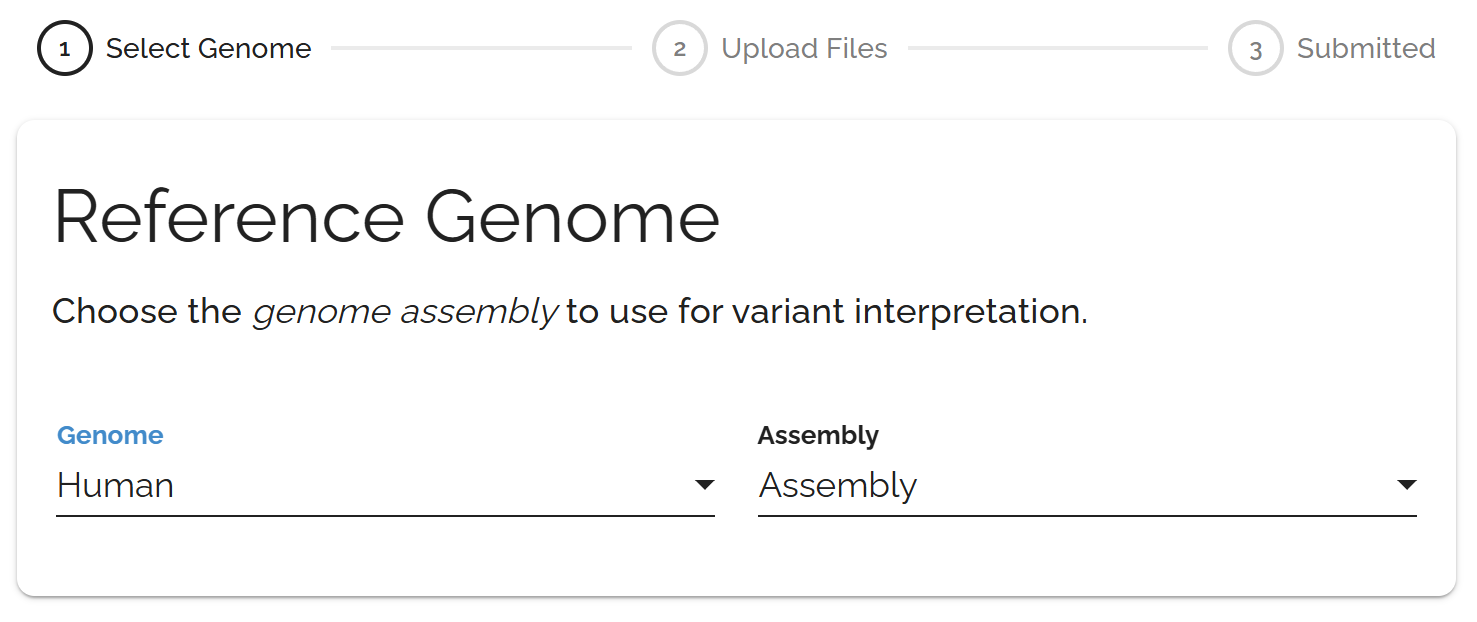
Upload your data
Upload genomic files (VCF/SNP format) and optionally add proteomics data (TMT or SomaScan). Bystro supports compressed files and can handle multiple files in a single submission.
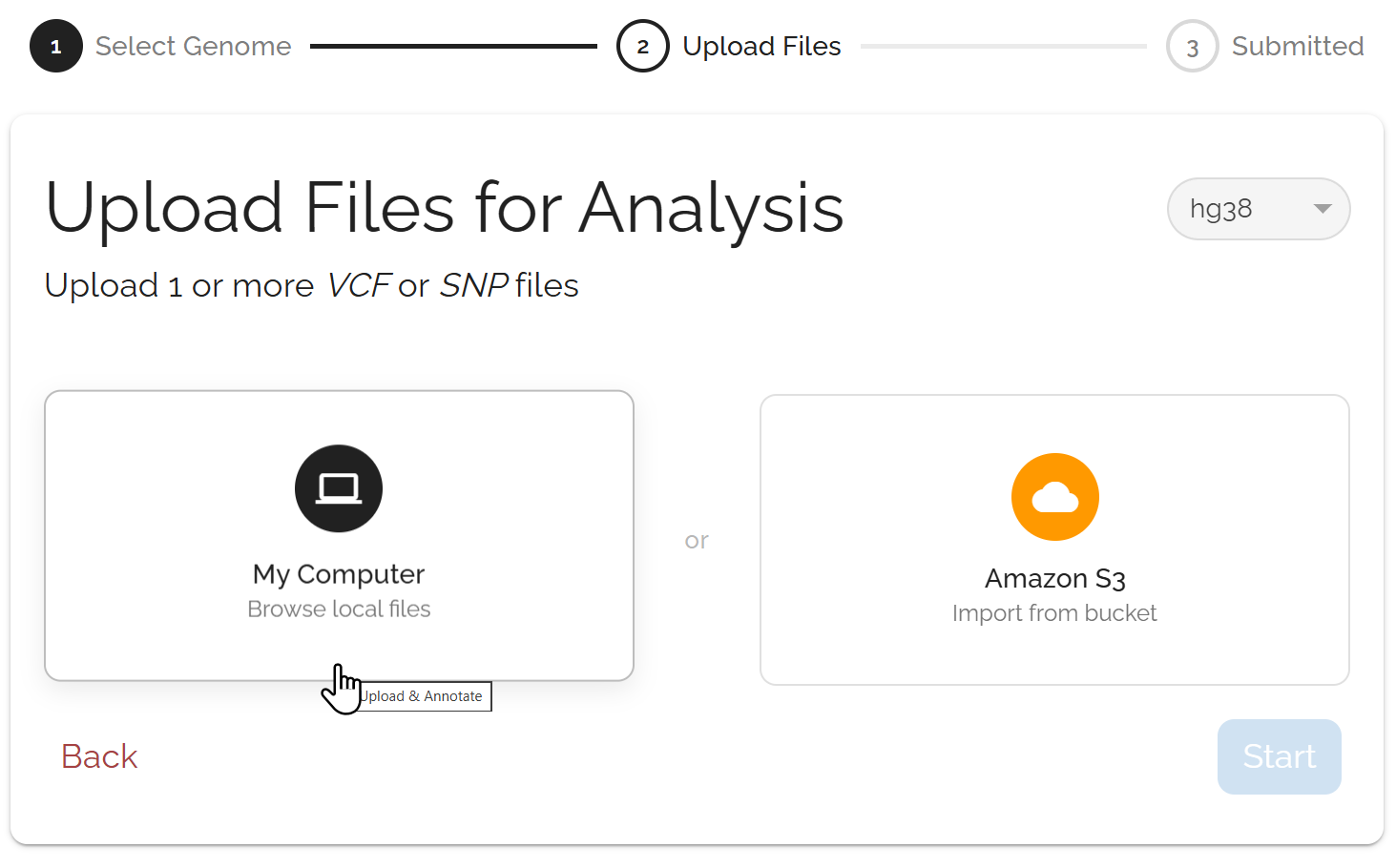
Configure advanced options
Optionally enable missing data imputation for SNP array data, add covariate information to link samples to phenotypes, or merge multiple chromosome files from the same subjects into a complete annotation.
Upload Methods
Desktop Upload
Upload files directly from your computer. Best for smaller files and simple workflows. Features include a drag-and-drop interface, progress tracking, and support for multiple files.
Amazon S3 Upload
Connect directly to your S3 buckets. Ideal for large files and cloud-based workflows. No file size limits, faster transfers for large datasets, and direct bucket browsing.
Advanced Features
Upload both genomic and proteomics data in a single submission to create integrated analyses.
Proteomics formats: TMT (Tandem Mass Tag) and SomaScan data.
Automatic linking: Bystro attempts to match samples between genomic and proteomic datasets.
Covariate integration: Shared covariates are automatically identified when possible.
Enhance SNP array datasets by imputing missing genotype data.
This feature fills in gaps in your genetic data using statistical methods, improving the completeness and analytical power of your dataset for downstream analyses.
Combine multiple chromosome files from the same subjects into one complete annotation.
Important: After selecting your chromosome files with checkboxes, you must click the Merge button. Simply checking the boxes does not initiate the merge.
Amazon S3 Integration
Set up your AWS credentials and upload files directly from your S3 storage.
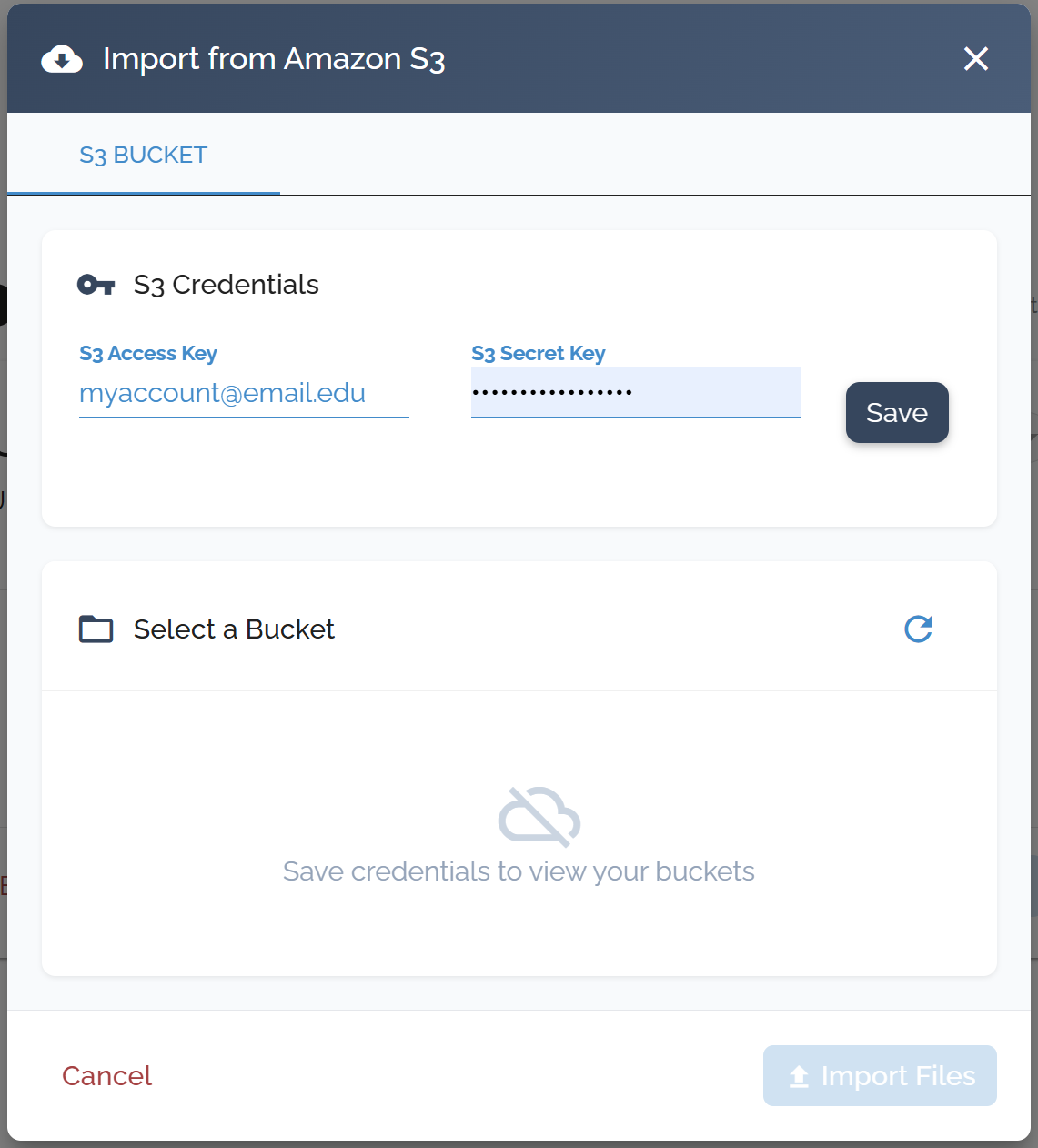
Supported File Formats
Genomic Data
VCF files (.vcf, .vcf.gz) or SNP format files (.snp), including WGS, WES, array data, and compressed formats. Multiple files per submission supported.
Proteomics Data
TMT (Tandem Mass Tags) and SomaScan proteomics datasets. Can be linked to genomic data during submission for integrated analysis.
Large files: Use S3 upload for files over 1GB for better reliability.
Multi-omics: Upload proteomics and genomics data together for integrated analysis.
Email notifications: Highly recommended for large datasets that take time to process.
Chromosome merging: Don't forget to click Merge after selecting your files.
Sample information: Add covariates during submission or afterward using the sample info feature.
What Happens Next
After submitting your files, Bystro will:
Validate your files
Check formatting and ensure your files meet the required standards.
Process advanced features
Run imputation, merging, or multi-omics integration if selected.
Queue your annotation
Place your job in the processing queue.
Annotate variants
Apply comprehensive annotations from all configured databases.
Generate integrated results
Make completed results available for download and analysis.
Accessing Your Genetic Files in THiNK
All annotated genetic files are available in THiNK. Click the DNA button in the bottom left of the chat box before asking a question to add your genetic data. Learn more in the THiNK overview.
